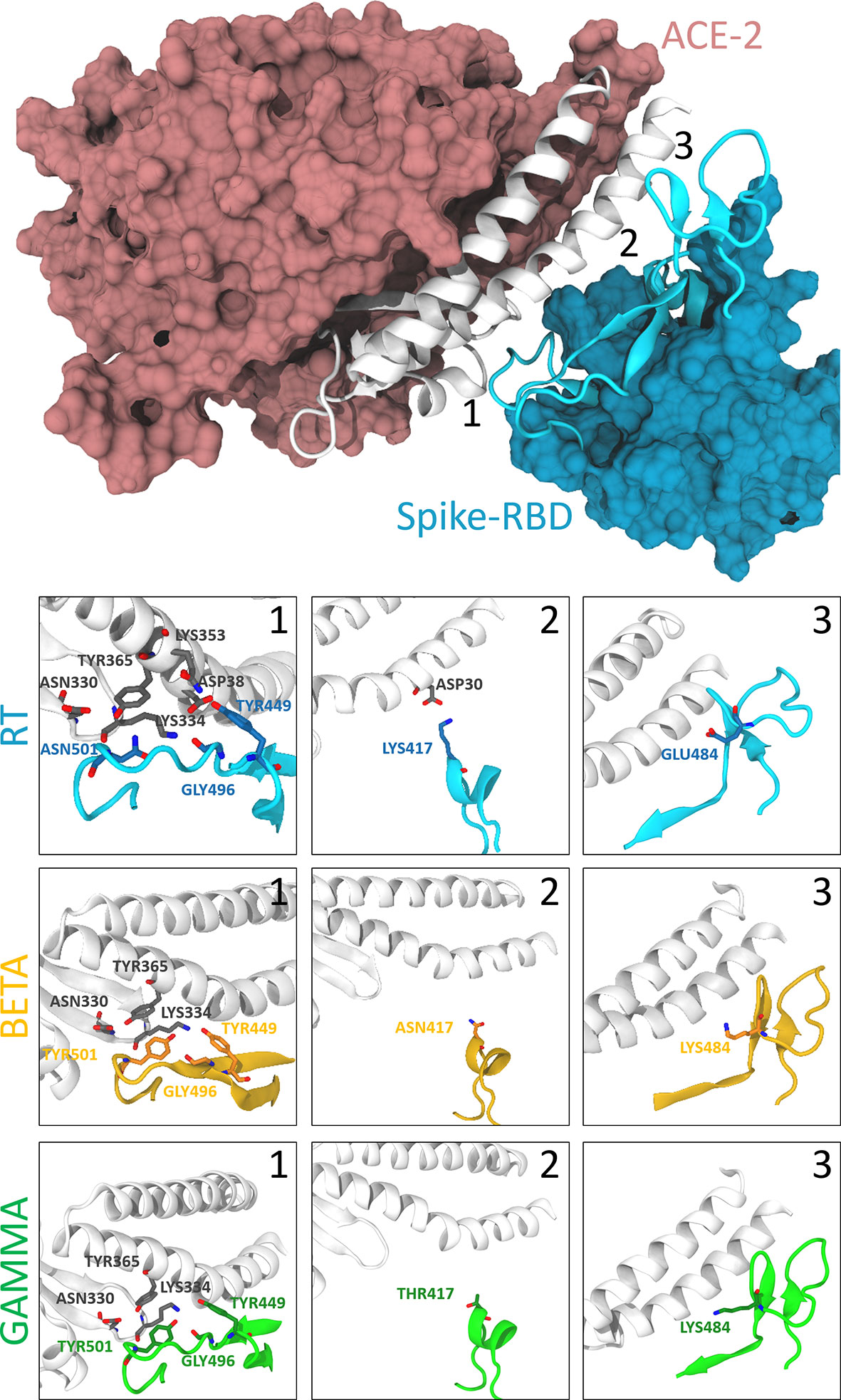
Molecular insights into receptor binding of recent emerging SARS-CoV-2 variants | Nature Communications
Frontiers | Rapid Assessment of Binding Affinity of SARS-COV-2 Spike Protein to the Human Angiotensin-Converting Enzyme 2 Receptor and to Neutralizing Biomolecules Based on Computer Simulations

On the binding affinity of macromolecular interactions: daring to ask why proteins interact | Journal of The Royal Society Interface

On the binding affinity of macromolecular interactions: daring to ask why proteins interact | Journal of The Royal Society Interface

Protein Binding Affinity of Polymeric Nanoparticles as a Direct Indicator of Their Pharmacokinetics | ACS Nano
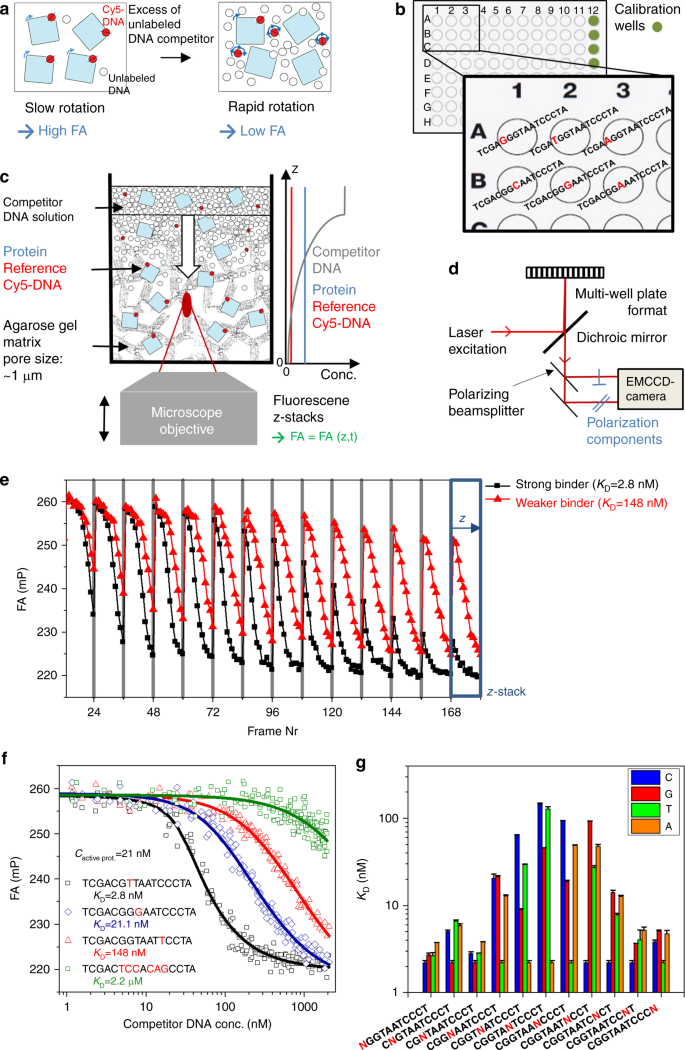
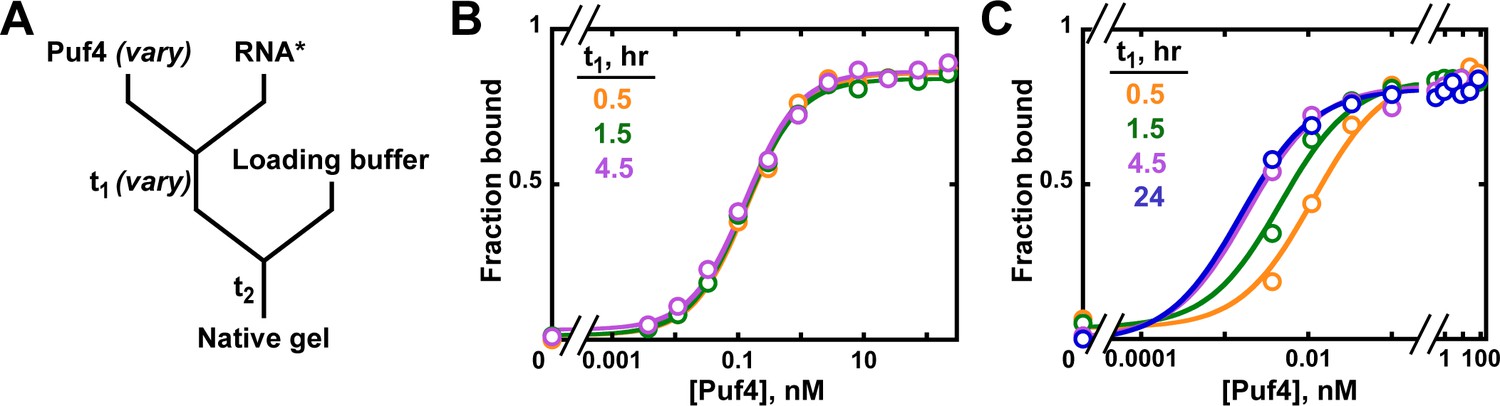
True equilibrium measurement of transcription factor-DNA binding affinities using automated polarization microscopy | Nature Communications
What determines a ligand's affinity for a receptor? Are there multiple factors in play that would make one ligand have a higher affinity than another for a given receptor? - Quora
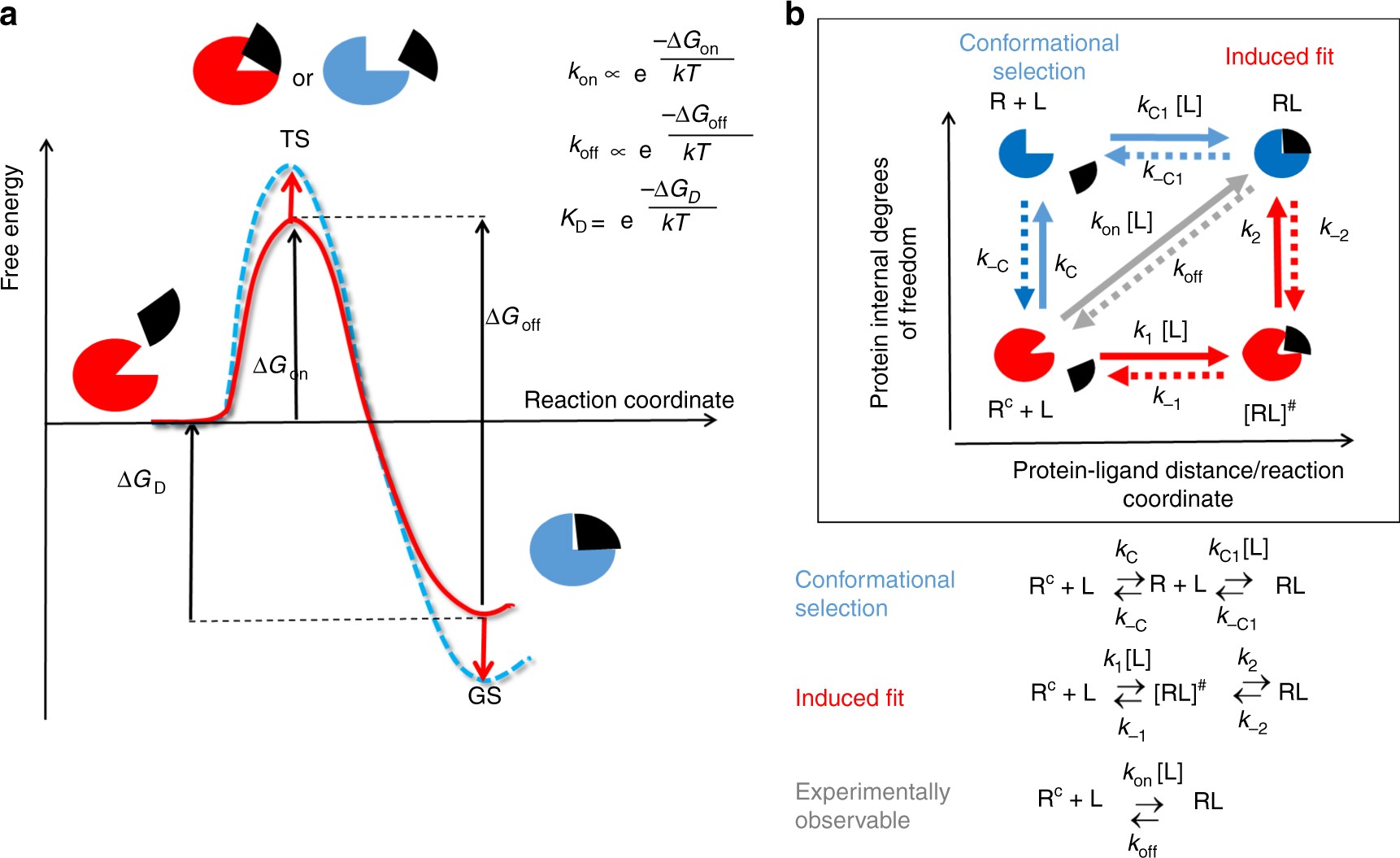
Protein conformational flexibility modulates kinetics and thermodynamics of drug binding | Nature Communications

In Situ Ligation of High‐ and Low‐Affinity Ligands to Cell Surface Receptors Enables Highly Selective Recognition - Taichi - 2017 - Advanced Science - Wiley Online Library
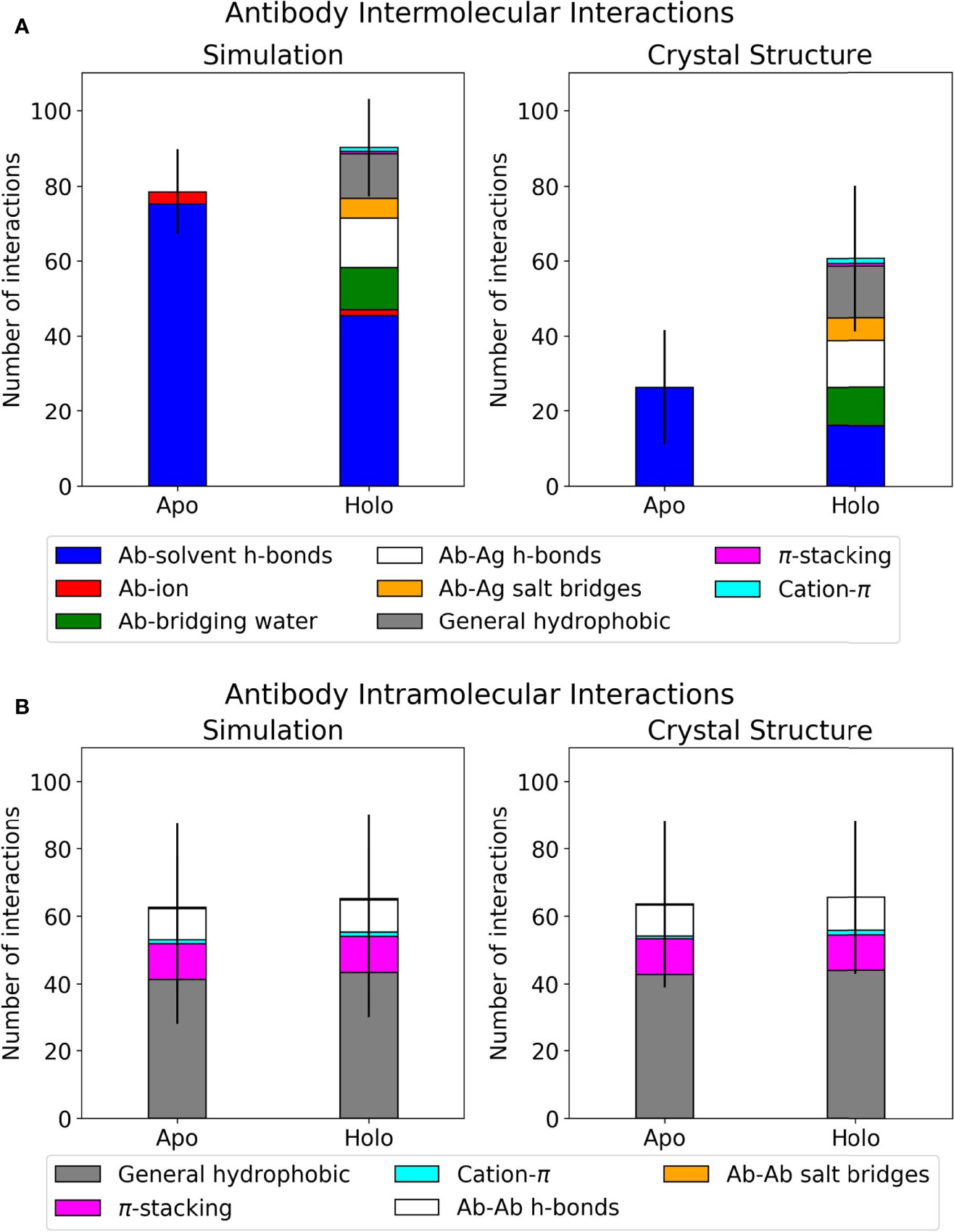
Frontiers | Higher Affinity Antibodies Bind With Lower Hydration and Flexibility in Large Scale Simulations
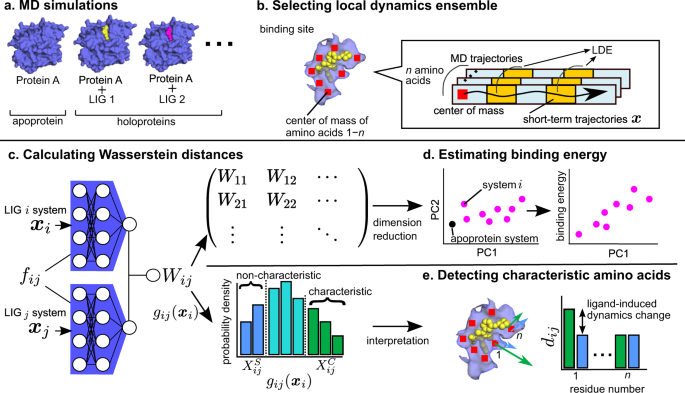
Differences in ligand-induced protein dynamics extracted from an unsupervised deep learning approach correlate with protein–ligand binding affinities | Communications Biology

Peptide Binder with High-Affinity for the SARS-CoV-2 Spike Receptor-Binding Domain | ACS Applied Materials & Interfaces