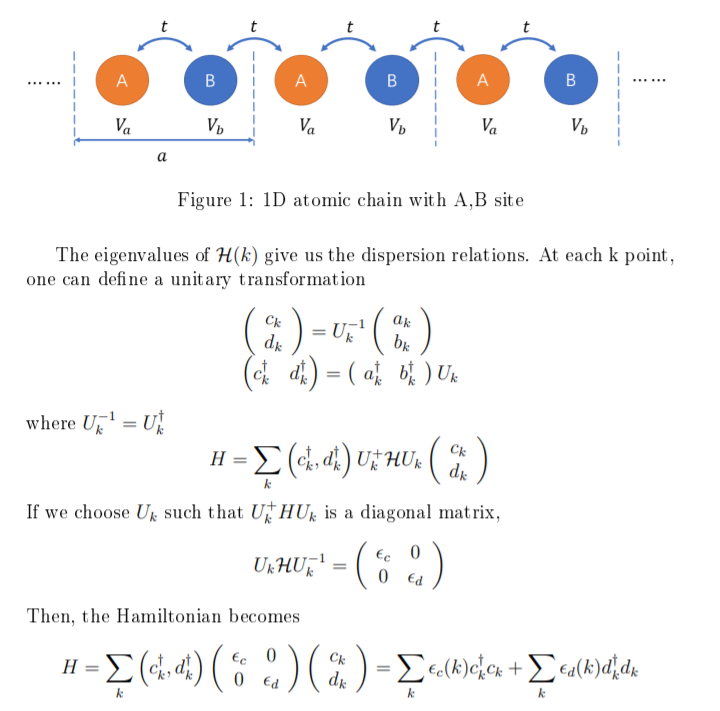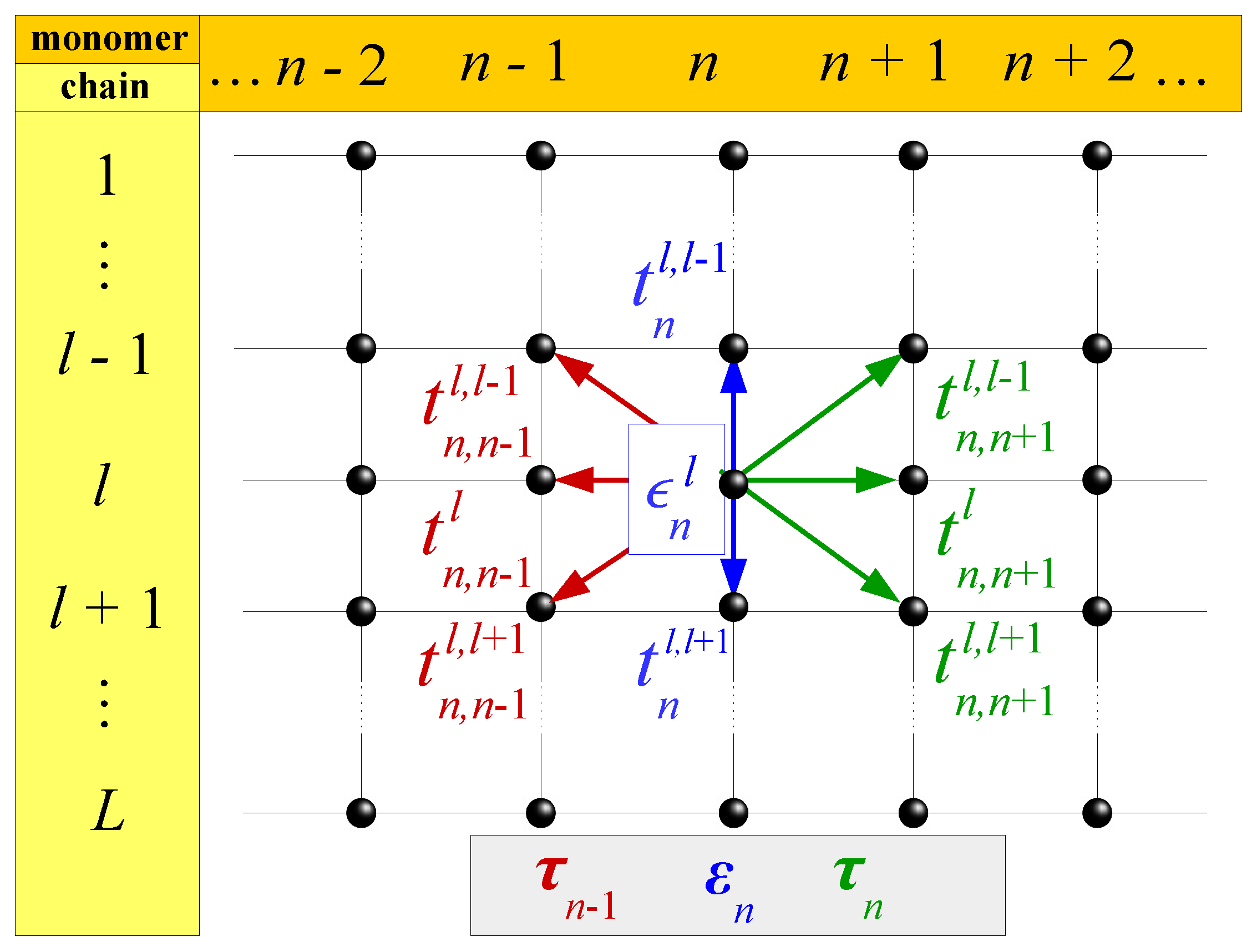
Symmetry | Free Full-Text | Tight-Binding Modeling of Nucleic Acid Sequences: Interplay between Various Types of Order or Disorder and Charge Transport

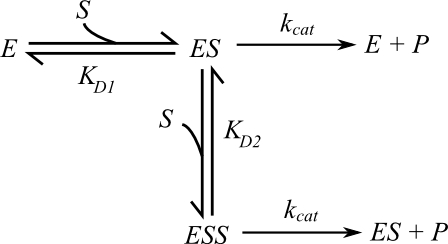
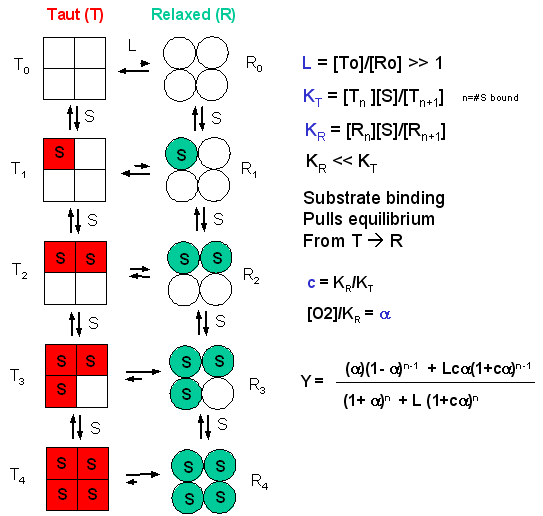
Binding models with two binding sites: Independent binding and negative... | Download Scientific Diagram

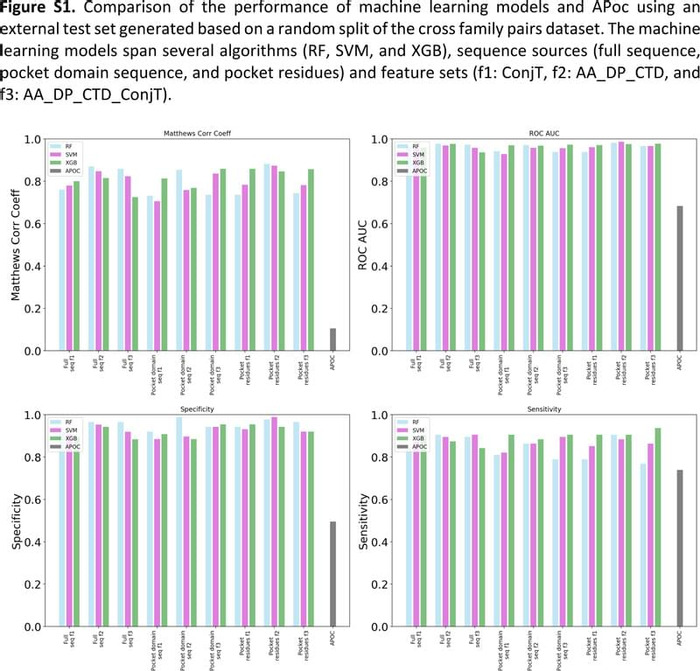
Conservation of binding properties in protein models - Computational and Structural Biotechnology Journal

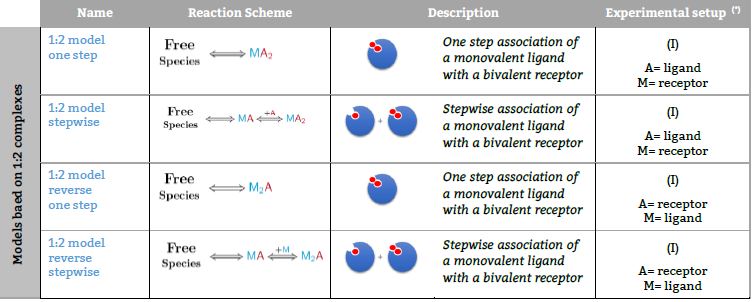
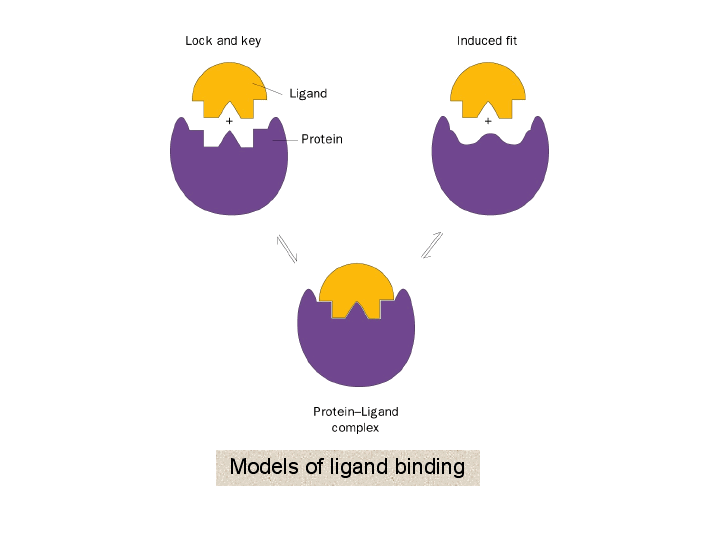
Schematic illustrations of the three protein-ligand binding models: (a)... | Download Scientific Diagram

Improving the Accuracy of Protein-Ligand Binding Affinity Prediction by Deep Learning Models: Benchmark and Model | Theoretical and Computational Chemistry | ChemRxiv | Cambridge Open Engage
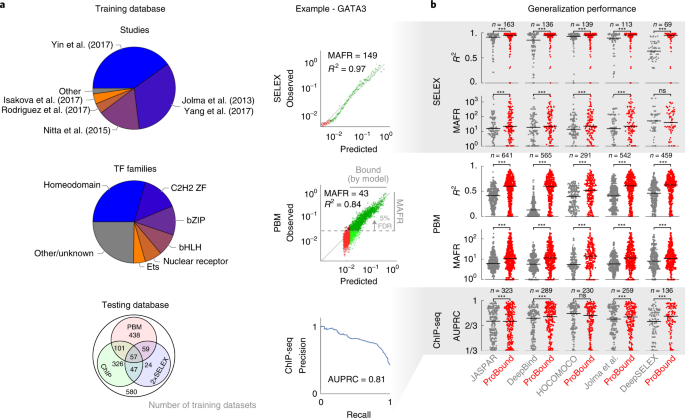
Prediction of protein–ligand binding affinity from sequencing data with interpretable machine learning | Nature Biotechnology